 Background and Artifact Removal for Multiplex Tissue Imaging
Background and Artifact Removal for Multiplex Tissue Imaging
Published:
I built this project to improve background and artifact removal in multiplex pathology images, including whole-slide imaging (WSI) and multi-channel microscopy data.
The core objective is to produce cleaner channels before downstream analysis steps such as:
- cell segmentation
- biomarker quantification
- spatial analysis
Why this matters
Multiplex tissue images often contain unwanted signals and artifacts, including:
- autofluorescence and residual staining noise
- non-tissue background
- pen/ink artifacts and bright splash-like outliers
If these remain in the data, biomarker signal estimates can be biased and scientific conclusions become less reliable. This is especially important in cancer-focused biomedical workflows where reproducibility and quantitative accuracy are critical.
Outcome
This workspace combines comparative methods and custom cleanup scripts into an applied preprocessing pipeline for multiplex tissue imaging. The output is cleaner channels and clearer overlays, improving reliability for downstream biomedical image analysis.
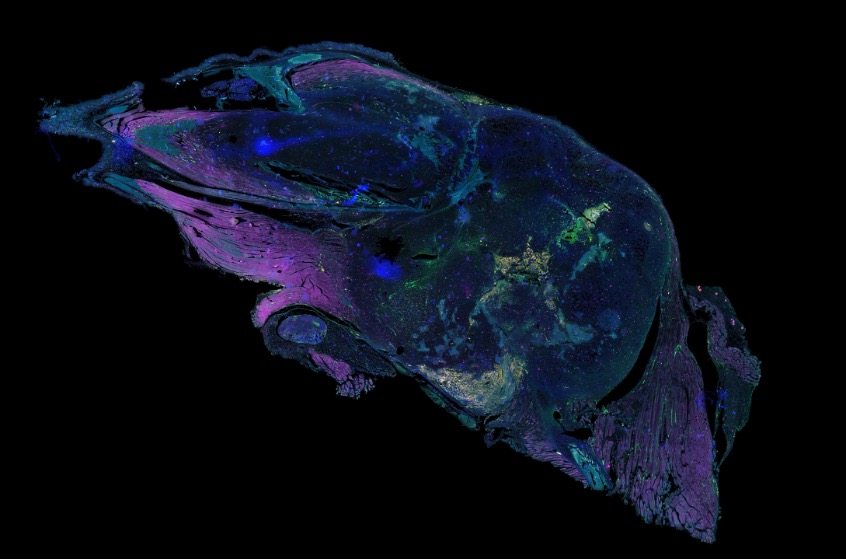
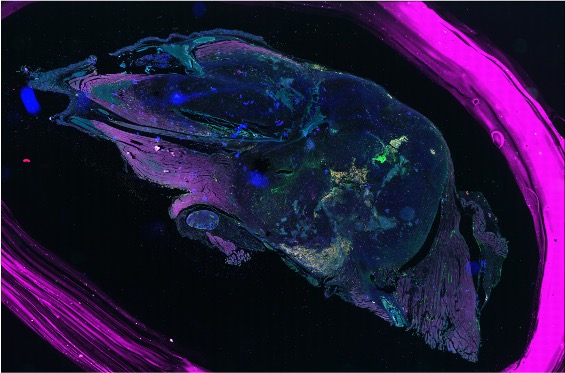
Before vs. after cleanup
Project repository: 0_Project: Background Removal for Multiplex Tissue Imaging